Human T-cell lymphotropic virus type III shares sequence homology with a family of pathogenic lentivirusesby MATTHEW A. GONDA, MICHAEL J. BRAUN, JANICE E. CLEMENTS, JOANNA M. PYPER, FLOSSIE WONG-STAAL, ROBERT C. GALLO, AND RAYMOND V. GILDEN
Proc. Natl. Acad. Sci. USA
Vol. 83, pp. 4007-4011, June 1986
NOTICE: THIS WORK MAY BE PROTECTED BY COPYRIGHTYOU ARE REQUIRED TO READ
THE COPYRIGHT NOTICE AT THIS LINK BEFORE YOU READ THE FOLLOWING WORK, THAT IS AVAILABLE SOLELY FOR PRIVATE STUDY, SCHOLARSHIP OR RESEARCH PURSUANT TO 17 U.S.C. SECTION 107 AND 108. IN THE EVENT THAT THE LIBRARY DETERMINES THAT UNLAWFUL COPYING OF THIS WORK HAS OCCURRED, THE LIBRARY HAS THE RIGHT TO BLOCK THE I.P. ADDRESS AT WHICH THE UNLAWFUL COPYING APPEARED TO HAVE OCCURRED. THANK YOU FOR RESPECTING THE RIGHTS OF COPYRIGHT OWNERS.
Human T-cell lymphotropic virus type III shares sequence homology with a family of pathogenic lentiviruses
(human and animal retroviruses/acquired immune deficiency syndrome/evolutionary relationships)
MATTHEW A. GONDA [ i], MICHAEL J. BRAUN [ i], JANICE E. CLEMENTS [ ii] , JOANNA M. PYPER [ ii], FLOSSIE WONG-STAAL [ oii], ROBERT C. GALLO [ iii], AND RAYMOND V. GILDEN [ iv]
ABSTRACT The etiologic agent of the acquired immune deficiency syndrome, human T-cell lymphotropic virus type m (HTLV-III), has recently been shown to morphologically resemble and share sequence homology with visna virus, a pathogenic lentivirus. Molecular hybridization, heteroduplex mapping, and DNA sequence analyses were used to compare HTLV-III to other lentiviruses of domestic animals, including visna, caprine arthritis encephalitis, and equine infectious anemia viruses. Hybridization results showed that a substantial amount of sequence homology exists between each of these viruses and HTLV-III. In addition, a closer relationship was found between visna and caprine arthritis encephalitis viruses than for any of the other lentiviruses studied. These results, along with nucleotide and amino acid sequence comparisons, have been used in a comprehensive effort to derive a systematic relationship for lentiviruses and to provide further evidence for classifying HTLV-III with the Lentivirinae subfamily of retroviruses. This relationship predicts that similarities in biology and disease process can be expected between HTLV-III and other Lentivirioae members.
Gonda et al. (1) recently showed that human T-cell lymphotropic virus type III (HTLV-III) morphologically resembles and shares nucleotide sequence homology with visna virus, a pathogenic, neurotropic lentivirus of sheep. Visna virus also shares other biological similarities with HTLV-III. For example, HTLV-III exhibits a strong tropism for cells of the immune system, in particular, helper T-lymphocytes upon which it exerts its biologic effects. In culture, HTLV-III induces syncytia and is cytolytic (2). Moreover, HTLV-III-induced acquired immune deficiency syndrome (AIDS) is characterized by T-cell depletion, immunosuppression, and opportunistic infections, and the propensity for HTLV-III to infect brain cells suggests a possible neuropathic role for HTLV-III in AIDS-related encephalopathy and dementia (2, 3). Visna virus also infects cells of the immune system (monocytes) and is cytolytic in cell culture, and it produces a slowly progressive inflammatory condition of the central nervous system, often leading to total paralysis and death from inanition (4-7).
Lentiviruses (subfamily Lentivirinae in the family Retroviridae) are exogenous, nononcogenic retroviruses (8) that cause persistent but debilitating infections, replicate at a slow but progressive rate, and have been shown to have pathogenic potential in vitro. Lentiviruses have been isolated from a variety of domestic ungulates including ovine (4-7), caprine (9-12), equine (13), and bovine (14) species.
The morphologic, genetic, and biologic similarities with visna virus provide strong evidence for a close evolutionary relationship between HTLV-III and the Lentivirinae subfamily of retroviruses (1). Further analysis of the evolutionary and systematic relationship between HTL V-III and other lentiviruses has awaited the molecular cloning of representative species (15-18). As a sequel to the initial findings of Gonda et al. (1), we have used heteroduplex mapping, molecular hybridization, and DNA sequence analyses to further detail the genetic relationship of HTL V-III, visna virus, caprine arthritis encephalitis virus (CAEV), and equine infectious anemia virus (EIAV) and to provide a phylogeny for this evolutionarily diverse family.
MATERIALS AND METHODSHeteroduplex Analysis. Heteroduplexes were prepared according to the method of Davis et at. (19) with modifications (1). The stringency of hybridization was calculated from the equations of McConaughy et al. (20) using the known G+C content (42%) of HTLV-III (21).
Southern Transfer Analysis. DNA fragments representing all or most of each lentiviral genome (see Fig. 2A) were electrophoresed, then transferred to nitrocellulose paper by the method of Southern (22). Nitrocellulose-bound DNAs were hybridized with the lentiviral probes (1 x 106 cpm/ml), which consisted of the virus genomes shown in Fig. 2A, radiolabeled with 32p to a specific activity of 1-3 x 108 cpm/ug by nick-translation (23). Hybridization was carried out in a buffer (24) containing 50% formamide and 1 M NaCI at 22°C [melting temperature (tm) - 40°C] for 72 hr. Filters were washed in 3x standard saline citrate (0.45 M NaCl/ 0.045 M sodium citrate, pH 7.0) at 37°C (tm - 55°C), and autoradiographed with Kodak XRP-5 x-ray film for 16 hr at -70°C with two Dupont Cronex Lightning Plus intensifying screens.
Nucleotide Sequencing. The visna virus sequence was determined from a previously described clone (16), by Maxam and Gilbert (25) chemical degradation from the BstEII site (see Fig. 3; nucleotides 71-77).
Phylogeny Reconstruction. Nucleotide sequences were aligned by the method of Dayhoff (26). The frequency of matching residues (M) was calculated from the optimal pairwise alignments, counting gaps as mismatches. By using -logM as a distance metric, a phylogenetic tree was con- structed by the method of Fitch and Margoliash (27) using the PHYLIP computer program package of J. Felsenstein (University of Washington, Seattle).
RESULTS AND DISCUSSIONWe first constructed heteroduplexes between HTLV-III and CAEV, a neurotropic virus of goats (9-12). CAE V shares a considerable amount of sequence homology with visna virus over its entire genome, but this homology is greatest for the gag and pol genes and for a highly conserved region overlapping the carboxyl terminus of pol, analogous to the sor (short open reading frame) region in HTLV-III (18,21,28).
Heteroduplexes between HTLV-III and CAE V spread at tm - 47°C revealed a substantial amount of homology (Fig. 1). Approximately 33% of the HTLV-III genome formed a duplex with CAEV. When the heteroduplexes were spread at tm - 39°C, 15% of the HTLV-III genome remained duplexed with CAE V , with the conserved sequences occurring largely in the gag/pol gene region (data not shown). This is in agreement with the extent and location of sequences conserved between HTLV-III and visna virus (1). As a negative control, we used a A clone ofHTLV-11I whose insert was in the reverse orientation of the clone used in Fig. 1A. No regions of hybridization were observed between CAEV and the reversed HTLV -III insert (data not shown), verifying that the conditions used for heteroduplex formation were not so mild as to allow nonspecific hybridization over short stretches of DNA. Furthermore, in previous heteroduplex studies between HTLV-III and HTLV-I or -II, conducted under similar low-stringency conditions, no hybridization was detected with HTLV-II and only a minimal amount was detected with HTL V-I (5-10% of that observed here between HTLV-III and CAEV) in the gag/pol region (1, 29).
 FIG. 1. Heteroduplexes were prepared with inserts in bacteriophage A. (A) Heteroduplex of HTLV-III and CAEV at tm - 47°C. (B) Interpretive drawing. The 5' and the 3' ends of the plus strand of the insert and the A arms are indicated. Regions of homology in A are represented by thick lines, and regions of nonhomology are represented by thin lines.
FIG. 1. Heteroduplexes were prepared with inserts in bacteriophage A. (A) Heteroduplex of HTLV-III and CAEV at tm - 47°C. (B) Interpretive drawing. The 5' and the 3' ends of the plus strand of the insert and the A arms are indicated. Regions of homology in A are represented by thick lines, and regions of nonhomology are represented by thin lines.To confirm and extend the heteroduplex data, we attempted to detect genetic relatedness among lentiviruses by using cloned viral genomes of HTLV -III, CAEV, EIA V, and visna virus (15-18) (Fig. 2A) in Southern transfer molecular hybridization experiments. The HTLV-III probe showed homology with all the genomes tested (Fig. 2B). No nonspecific hybridization was detected with an equimolar amount of negative control DNA. In reciprocal hybridizations, EIA V and visna virus probes also detected HTLV-III and other lentiviruses (Fig. 2B). As the washing stringencies were increased (1m -34°C to -25°C), the heterologous hybridization intensities were reduced while the homologous reaction remained undiminished. Furthermore, when purified inserts were molecularly dissected with appropriate restriction enzymes, hybridization experiments showed that the greatest amount of homology between each of these viruses exists in the pol or the gag/pol region (data not shown) as has been shown for HTLV-III and visna virus (1).
To help quantify the relationship between HTLV-III, lentiviruses, and other retroviruses, we determined a ml\ior portion of the nucleotide sequence for a visna virus clone. A segment of this sequence from a region near the amino terminus of the visna virus pol gene is shown in Fig. 3. The nucleotide sequence homology between visna virus and HTL V-III is 66% over this entire region. Several segments of 25-40 nucleotides in this region have greater than 80% homology (visna nucleotides 1-39, 109-142, and 232-255), and several stretches have from 8 to 13 bases identical. Clearly, this degree of sequence conservation accounts for the specific hybridization seen under the relaxed conditions of the heteroduplex and Southern blot analyses (Figs. 1 and 2; refs. 1, 29) and contrasts strongly with the negative hybridization results between cloned visna virus and LA V reported by others (30, 31). An independently determined visna virus sequence from a different clone matches our sequence (Fig. 3) except for a single silent substitution at nucleotide 78 and shows a similar degree of conservation over the rest of the pol gene (28), thus further confirming our initial observations on the genetic relationship of visna virus and HTLV-III (1).
The predicted amino acid sequences for HTLV -III and the visna virus pol gene segments shown in Fig. 3 were aligned with the pol sequence of EIA V (17), CAEV (32), and several other representative retroviruses (Fig. 4A). It is obvious from inspection of this alignment that HTLV-III, visna virus, CAEV, and EIA V cluster together. Using the amount of amino acid homology computed for each pairwise comparison, we next derived a phylogenetic tree for these viral genes (Fig. 4B). In rooting this tree, Moloney murine leukemia virus (Mo-MuLV) was considered to be the outgroup taxon because it consistently had the lowest alignment score with each of the other viruses. This decision is supported by the fact that the reverse transcriptase of Mo-MuLV preferentially uses Mn2+ as cofactor, while the reverse transcriptases of all the other viruses prefer Mg2+, and by previous artalyses of the relationships of retroviral pol genes, which identified the mammalian type C pol genes as the most divergent (35).
The phylogenetic tree (Fig. 4B) clearly shows that HTL VIII clusters with the lentiviruses. Within this group, visna virus and CAEV are the most closely related with amino acid homology of 90%; HTL V -III is slightly closer to this pairthan is EIA V. The amino acid homology between HTL V -III, EIA V, and the CAEV /visna virus pair is approximately 60%. The next closest viruses are Rous sarcoma virus (RSV) and HTLV-I at about 40% amino acid homology. Finally, MoMuL V was the most divergent, with about 30% amino acid homology to the lentiviruses. While the tree topology derived from this region of the pol gene has HTL V-III more closely related to visna virus and CAEV than either is to EIAV, sequence comparisons from other segments of the genome suggest alternative tree topologies with HTLV-III or CAEV /visna virus diverging first (17). Therefore, aside from the close relationship of visna virus and CAEV, we consider the lentiviruses to be about equally related. Overall, the tree topology is in general agreement with other recent attempts at classification of retroviral pol genes (28, 32).
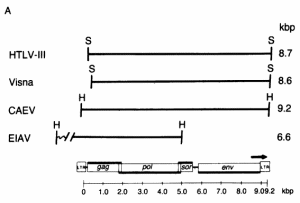 FIG. 2. Detection of sequence homology between HTL V-III, CAEV, EIAV, and visna virus by molecular hybridization. (A) Restriction enzyme fragments used in the hybridization studies. The EIAV fragment contained about 1.4 kilobase pairs (kbp) of host flanking sequences at its left end. The relative genomic organization of each fragment is indicated by its alignment with the generalized lentivirus gene map shown at the bottom. The arrow over the env gene region indicates the 3' open reading frame (ORF) in HTLV-Ill. The genomic organization was determined from the nucleotide sequence for HTLV-III (21), EIAV (17), and visna virus (28) and by heteroduplex mapping for CAEV (18). H, HindIII; S, Sac I.
FIG. 2. Detection of sequence homology between HTL V-III, CAEV, EIAV, and visna virus by molecular hybridization. (A) Restriction enzyme fragments used in the hybridization studies. The EIAV fragment contained about 1.4 kilobase pairs (kbp) of host flanking sequences at its left end. The relative genomic organization of each fragment is indicated by its alignment with the generalized lentivirus gene map shown at the bottom. The arrow over the env gene region indicates the 3' open reading frame (ORF) in HTLV-Ill. The genomic organization was determined from the nucleotide sequence for HTLV-III (21), EIAV (17), and visna virus (28) and by heteroduplex mapping for CAEV (18). H, HindIII; S, Sac I. 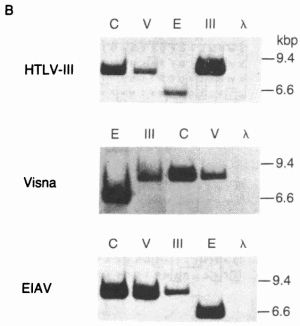 (B) Southern transfers of the lentiviral genomic fragments shown in A hybridized with 32P-labeled HTLV-III, visna, and EIA V probes under relaxed conditions (tm - 40°C). The amount of DNA loaded into each lane was 100 pg for lanes homologous to the probe and 100 ng for heterologous lanes, except for the hybridization of CAEV with the visna probe, for which 1 ng of CAEV DNA was used because of the extensive homology between CAEV and visna. Each gel also contained 500 ng of HindIII-digested DNA fragments as molecular weight markers and negative hybridization controls. Lanes: C, CAEV; V, visna; E, EIAV; III, HTLV-III; A.
(B) Southern transfers of the lentiviral genomic fragments shown in A hybridized with 32P-labeled HTLV-III, visna, and EIA V probes under relaxed conditions (tm - 40°C). The amount of DNA loaded into each lane was 100 pg for lanes homologous to the probe and 100 ng for heterologous lanes, except for the hybridization of CAEV with the visna probe, for which 1 ng of CAEV DNA was used because of the extensive homology between CAEV and visna. Each gel also contained 500 ng of HindIII-digested DNA fragments as molecular weight markers and negative hybridization controls. Lanes: C, CAEV; V, visna; E, EIAV; III, HTLV-III; A.Because the amino acid sequences provided a data set of discrete characters including a well-defined outgroup (MoMuLV), we used cladistic analysis (36) to confirm the kinship of HTLV-III and lentiviruses. This analysis revealed that there are at least 10 positions where visna virus, CAEV, HTLV -III, and EIAV share a potentially derived character state (shared amino acids that are not shared with the other viruses). Of these, residues 11, 28, 43, 46, and 59 (Fig. 4A) are especially good candidates, because HTLV-I, RSV, or both share the putative ancestral character state with Mo-MuL V while visna virus, CAEV, HTLV-III, and EIAV share a potentially derived character state. Thus, beyond their overall similarity, HTL V-III is united with the lentivirus group by a number of probable shared derived characters.
Our efforts to detect and compare conserved nucleotide and amino acid sequences in the pol gene of lentiviruses have enabled us to derive the first comprehensive systematic relationship for this group. Biologically significant similarities extend to the genomic organization of these viruses. For example, the topographies of the HTLV -III and visna virus genomes, like those of other retroviruses, contain gag, pol, and env structural genes located 5' to 3', respectively. In addition, there are two extra ORFs in HTLV-III that are unique among retroviruses (21, 30, 34, 37-39). The first ORF, sor, is located immediately 3' to the pol gene and partially overlaps its carboxyl terminus. The second (3' ORF) overlaps the carboxyl terminus of the env gene and extends into the 3' long terminal repeat (LTR) (21,30,39). Visna virus (28) and CAEV, based on the extensive sequence homology of CAE V with visna virus in this region (18), contain a sor. A 3' ORF has not been identified in the one visna virus clone sequenced to date (28).
A novel feature of HTL V -III and visna virus is a gene called tat, for trans-activating transcriptional regulation, that encodes a factor that is believed to modulate viral expression in infected cells (40, 41). HTLV-I and -II and bovine leukemia virus also contain a tat gene, which has been localized to a region between the env gene and the 3' LTR (42-44), called pX (33). However, in contrast to these viruses, the tat gene for HTLV-III is not contained in the analogous 3' post-en v region (3' ORF) but is in the middle of the genome in a region between the sor and the env genes (41). Such a region also exists for visna virus (28) and is strongly conserved both structurally and functionally between visna virus and CAEV (18, 40), which, by extrapolation, suggests that the tat gene for both of these viruses will also be localized to this area.
 FIG. 3. Nucleotide sequence comparison between homologous regions of the pol genes of visna virus and HTLV-III. The HTLV-III sequence is from the BH10 clone (21); published sequences of lymphadenopathy virus (LAV) and HTLV-III are identical in this region (30).
FIG. 3. Nucleotide sequence comparison between homologous regions of the pol genes of visna virus and HTLV-III. The HTLV-III sequence is from the BH10 clone (21); published sequences of lymphadenopathy virus (LAV) and HTLV-III are identical in this region (30).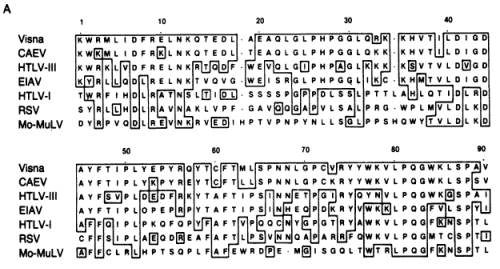 FIG. 4. (A) Alignment of lentivirus pol genes with those of other retroviruses. The visna virus amino acid sequence predicted by the segment of nucleotide sequence shown in Fig. 3 was aligned with those of six other retrovirus pol genes including HTLV-III (21), EIAV (17), CAEV (32), HTLV-I (33), RSV (34), and Mo-MuLV (30). The alignment shown for each virus is generally that optimal with visna virus; slight improvements in the other pairwise alignment scores can be made by minor shifts in the placement of gaps. Boxes are drawn around identical residues when at least two lentiviruses (visna virus, CAEV, HTLV-III, or EIAV) share that residue.
FIG. 4. (A) Alignment of lentivirus pol genes with those of other retroviruses. The visna virus amino acid sequence predicted by the segment of nucleotide sequence shown in Fig. 3 was aligned with those of six other retrovirus pol genes including HTLV-III (21), EIAV (17), CAEV (32), HTLV-I (33), RSV (34), and Mo-MuLV (30). The alignment shown for each virus is generally that optimal with visna virus; slight improvements in the other pairwise alignment scores can be made by minor shifts in the placement of gaps. Boxes are drawn around identical residues when at least two lentiviruses (visna virus, CAEV, HTLV-III, or EIAV) share that residue. 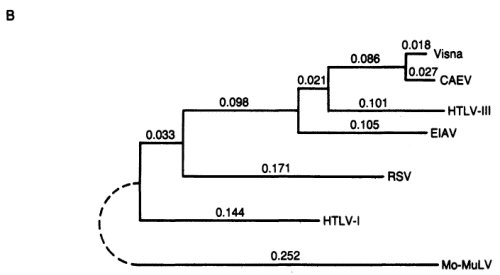 (B) Fitch-Margoliash phylogenetic tree of retroviral relationships based on the pol gene region shown in A. Branch lengths are in units of -logM, where M is the frequency of matching residues. The tree was rooted with Mo-MuL V as the outgroup. The average percent standard deviation of the tree was 3.3.
(B) Fitch-Margoliash phylogenetic tree of retroviral relationships based on the pol gene region shown in A. Branch lengths are in units of -logM, where M is the frequency of matching residues. The tree was rooted with Mo-MuL V as the outgroup. The average percent standard deviation of the tree was 3.3.HTLV-III shares other structural and biologic features in common with lentiviruses. On the basis of their morphologic fine structure, the viruses now classified as Lentivirinae are indistinguishable from HTLV-III (45-47). They all contain, in their outer membrane, a very large (90-135 kDa) glycosylated envelope protein and a transmembrane protein of 40 kDa (48-52); the large size of the envelope protein is unique to this subfamily of retroviruses. The major core protein (P24) of LAV and HTLV-III can be immunoprecipitated by antibodies to EIAV (53), predicting that nucleotide sequence homology would be found in the gag gene as recently demonstrated (17).
HTLV-III is a lentivirus as supported by the criteria outlined above. However, there are indications that there may be other family members more closely related to HTLV-III than those we have analyzed. A new viral isolate, simian T-lymphotropic virus type III (STLV-III), possesses ultrastructural morphology and other biologic properties similar to those of lentiviruses and viral proteins that are immunologically crossreactive with those of HTLV-III (54, 55). These data imply a genetic relationship for HTLV-III and STLV-III. If such is the case, it will be important to isolate molecular clones to establish the relationship between STL V-III and the lentiviruses used in this study.
Many factors in lentivirus-host interactions may have relevance to HTLV -III-induced disease. CAEV (56), EIAV (57), and bovine visna-like virus (BVV) (14), like HTLV-III and visna virus, also infect the cells of the immune system and, although the clinical sequelae vary for each virus, parallels with AIDS are apparent. Visna infection is associated with chronic pneumonitis, encephalitis, and wasting (6, 7); CAEV with crippling arthritis and encephalitis (9-12); EIAV with intermittent anemia, bouts of fever, and immune-complex glomerulonephritis (13); and BYV with lymphadenopathy and persistent lymphocytosis (14).
Despite differences in the outcome with infection by lentiviruses, one point remains in common; infected individuals remain infected for life. The exact mechanism of persistence by HTLV-III is not known. Lentiviruses, like other retroviruses, can integrate into the host cell DNA and produce latent infections. Once expressed, lentiviruses have developed different strategies to evade elimination by the immune response of the host. For example, visna virus and EIA V induce binding antibodies to all of their respective polypeptides and titratable neutralizing antibodies; however, "antigenic drift" in their env genes yields variants that escape immune surveillance mechanisms and induce a new cycle of disease (58, 59). CAEV also induces binding antibodies but, in contrast, titratable neutralizing antibodies are rare, and they are effective against only a narrow range of CAEV strains (60).
Genomic diversity for HTLV-III, particularly in the env region, is well documented (21,39,61,62) and is suggestive of antigenic variation. Neutralizing antibodies of low titer have also been detected in patients with AIDS and AIDS-related complex (63, 64). However, in one of these studies (64), neither early nor late serum relative to HTLV-III isolation from a single individual was strongly neutralizing, and different viral isolates from geographically separated areas did not vary significantly in sensitivity to neutralization by selected sera. Thus, the mechanism of persistence for HTLV-III may be more reminiscent of CAEV in that neutralizing antibodies are rarely achieved. The phylogenetic relationship of HTLV-III and lentiviruses imply that a better understanding of mechanisms of lentivirus persistence and the parameters of neutralization by antibodies could contribute t9 the effective control of HTLV-III-induced disease.
We thank N. Rice for allowing use of her EIAV nucleotide sequence data; K. Nagashima, J. Shumaker, and J. E. Elser for technical contributions; D. Lomb for assistance with computer analysis; J. Felsenstein for advice on the use of PHYLIP; and J. Hopkins for assistance in preparing the manuscript. M.A.G., M.J .B., and R.V.G. were supported by Public Health Service Contract NOl-CO-23910 ·with Program Resources, Inc., from the National Cancer Institute.
_______________
Notes:1. Gonda, M. A., Wong-Staal, F., Gallo, R. C., Clements, J. E., Narayan, O. & Gilden, R. V. (1985) Science 227, 173-177.
2. Wong-Staal, F. & Gallo, R. C. (1985) Nature (London) 317, 395-403.
3. Shaw, G. M., Harper, M. E., Hahn, B. H., Epstein, L. G., Gajdusek, D. C., Price, R. W., Navia, B. A., Petito, C. K., O'Hara, C. J., Groopman, J. E., Cho, E.-S., Oleske, J. M., Wong-Staal, F. & Gallo, R. C. (1985) Science 227, 177-182.
4. Narayan, 0., Wolinsky, J. S., Clements, J. E., Strandberg, J. D., Griffin, D. E. & Cork, L. C. (1982) J. Gen. Virol. 59, 345-356.
5. Haase, A. T. (1975) Curr. Top. Microbiol. Immunol. 72, 101-156.
6. Sigurdsson, B. (1954) Brit. Vet. J. 110, 225-270.
7. Sigurdsson, B., Palsson, P. A. & Grimsson, H. (1957) J. Neuropathol. Exp. Neurol. 16, 389-403.
8. Matthews, R. E. (1979) Intervirology 12, 234-238.
9. Crawford, T. B., Adams, D. S., Cheevers, W. P. & Cork, L. C. (1980) Science 207, 997-999.
10. Adams, D. S., Crawford, T. B. & Klevjer-Anderson, P. (1980) Am. J. Pathol. 99, 257-278.
11. Cork, L. C., Hadlow, W. J., Crawford, T. B., Gorham, J. R. & Piper, R. C. (1974) J. Infect. Dis. 129, 134-141.
12. Cork, L. C. & Narayan, O. (1980) Lab. Invest. 42, 596-602.
13. Crawford, T. B., Cheevers, W. P., Klevjer-Anderson, P. & McGuire, T. C. (1978) in Persistent Viruses, ICN/UCLA Symposium on Molecular and Cellular Biology, eds. Stevens, J., Todaro, G. & Fox, C. F. (Academic, New York), Vol. 2, pp. 727-749.
14. Van Der Maaten, M. J., Boothe, A. D. & Seger, C. L. (1972) J. Natl. Cancer Inst. 49, 1649-1657.
15. Hahn, B. H., Shaw, G. M., Arya, S. K., Popovic, M., Gallo, R. C. & Wong-Staal, F. (1984) Nature (London) 312, 166-169.
16. Molineaux, S. & Clements, J. E. (1983) Gene 23, 137-148.
17. Stephens, R. M., Casey, J. W. & Rice, N. R. (1985) Science 231, 589-594.
18. Pyper, J. M., Clements, J. E. & Gonda, M. A. (1985) J. Virol., in press.
19. Davis, R. W., Simon, M. & Davidson, N. (1971) Methods Enzymol. 21, 413-428.
20. McConaughy, B. L., Laird, C. D. & McCarthy, B. J. (1969) Biochemistry 8, 3289-3295.
21. Ratner, L., Haseltine, W., Patarca, R., Livak, K. J., Starcich, B., Josephs, S. J., Doran, E. R., Antoni Rafalski, J., Whitehorn, E. A., Baumeister, K., Ivanoff, L., Petteway, S. R., Jr., Pearson, M. L., Lautenberger, J. A., Papas, T. S., Ghrayeb, J., Chang, N. T., Gallo, R. C. & Wong-Staal, F. (1985) Nature (London) 313, 277-284.
22. Southern, E. M. (1975) J. Mol. Bioi. 98, 503-517.
23. Rigby, P. W. J., Dieckmann, M., Rhodes, C. & Berg, P. (1977) J. Mol. Biol. 113, 237-251.
24. Mullins, J. I., Casey, J. W., Nicolson, M. O. & Davidson, N. (1980) Nucleic Acids Res. 8, 3287-3305.
25. Maxam, A. M. & Gilbert, W. (1980) Methods Enzymol. 65, 499-560.
26. Dayhoff, M. O. (1978) in Atlas of Protein Sequence and Structure, ed. Dayhoff, M. O. (Natl. Biomedical Research Found., Washington, DC), Vol. 5, Suppl. 3, pp. 1-8.
27. Fitch, W. M. & Margoliash, E. (1967) Science 155, 279-284.
28. Sonigo, P., Alizon, M., Staskus, K., K1atzmann, D., Cole, S., Danos, 0., Retzel, E., Tiollais, P., Haase, A. & Wain-Hobson, S. (1985) Cell 42, 369-382.
29. Gonda, M. A., Wong-Staal, F., Gallo, R. C., Clements, J. C. & Gilden, R. V. (1985) Cancer Res. 45, 4553-4558.
30. Alizon, M., Sonigo, P., Barre-Sinoussi, F., Chermann, J.-C., Tiollais, P., Montagnier, L. & Wain-Hobson, S. (1984) Nature (London) 312, 757-760.
31. Wain-Hobson, S., Alizon, M. & Montagnier, L. (1985) Nature (London) 313, 743.
32. Chiu, I.-M., Yaniv, A., Dalberg, J. E., Gazit, A., Skuntz, S. F., Tronick, S. R. & Aaronson, S. A. (1985) Nature (London) 317, 366-368.
33. Seiki, M., Hattori, S., Hirayama, Y. & Yoshida, M. (1983) Proc. Natl. Acad. Sci. USA SO, 3618-3622. Proc. Natl. Acad. Sci. USA 83 (1986) 4011
34. Teich, N. (1984) in Molecular Biology of Tumor Viruses: RNA Tumor Viruses, eds. Weiss, R. A., Teich, N., Varmus, H. & Coffin, J. (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY), pp. 25-208.
35. Chiu, I.-M., Callahan, R., Tronick, S. R., Schlom, J. & Aaronson, S. A. (1984) Science 223, 364-370.
36. Wiley, E. O. (1984) Phylogenetics: The Theory and Practice of Phylogenetic Systematics (Wiley, New York).
37. Rice, N. R., Stephens, R. M., Couez, D., Deschamps, J., Kettmann, R., Burny, A. & Gilden, R. V. (1984) Virology 138, 82-93.
38. Rice, N. R., Stephens, R. M., Burny, A. & Gilden, R. V. (1985) Virology 142, 357-377.
39. Sanchez-Pescador, R., Power, M. D., Barr, P. J., Steimer, K. S., Stempien, M. M., Brown-Shimer, S. L., Gee, W. W., Renard, A., Randolph, A., Levy, J. A., Dina, D. & Luciw, P. A. (1985) Science 227, 484-492.
40. Hess, J. L., Clements, J. E. & Narayan, O. (1985) Science 229, 482-485.
41. Sodroski, J., Patarca, R., Rosen, C., Wong-Staal, F. & Haseltine, W. (1985) Science 229, 74-77.
42. Derse, D., Caradonna, S. J. & Casey, J. W. (1985) Science 227, 317-320.
43. Sodroski, J., Rosen, C., Wong-Staal, F., Salahuddin, S. Z., Popovic, M., Arya, S., Gallo, R. C. & Haseltine, W. A. (1985) Science 227, 171-173.
44. Sodroski, 1., Rosen, C., Goh, W. C. & Haseltine, W. (1985) Science 228, 1430-1434.
45. Thormar, H. (1961) Virology 14, 463-475.
46. Coward, J. E., Harter, D. H. & Morgan, C. (1970) Virology 40, 1030-1038.
47. Gonda, M. A., Charman, H. P., Walker, J. L. & Coggins, L. (1978) Am. J. Vet. Res. 39, 731-740.
48. Vigne, R., Filippi, P., Querat, G., Sauze, N., Vitu, C., Russo, P. & Delori, P. (1982) J. Virol. 42, 1046-1056.
49. Pyper, J. M., Clements, J. E., Molineaux, S. M. & Narayan, O. (1984) J. Virol. 51, 713-717.
50. Payne, S., Parekh, B., Montelaro, R. C. & Issei, C. J. (1984)J. Gen. Virol. 65, 1395-1399.
51. Robey, W. G., Safai, B., Oroszlan, S., Arthur, L. 0., Gonda, M. A., Gallo, R. C. & Fischinger, P. J. (1985) Science 228, 593-595.
52. Veronese, F. D., DeVico, A. L., Copeland, T. D., Oroszlan, S., Gallo, R. C. & Samgadharan, M. G. (1985) Science 229, 1402-1405.
53. Casey, J. M., Kim, Y., Andersen, P. R., Watson, K. F., Fox, J. L. & Devare, S. G. (1985) J. Virol. 55, 417-423.
54. Kanki, P. J., McLane, M. F., King, N. W., Jr., Letvin, N. L., Hunt, R. D., Sehgal, P., Daniel, M. D., Desrosiers, R. C. & Essex, M. (1985) Science 228, 1199-1201.
55. Daniel, M. D., Letvin, N. L., King, N. W., Kannagi, M., Sehgal, P. K., Hunt, R. D., Kanki, P. J., Essex, M. & Desrosiers, R. C. (1985) Science 228, 1201-1204.
56. Narayan, 0., Kennedy-Stoskopf, S., Sheffer, D., Griffin, D. E. & Clements, J. E. (1983) Infect. Immun. 41, 67-73.
57. Kobayashi, K. & Kono, Y. (1967) Nat. Inst. Anim. Health Q. 7, 8-20.
58. Narayan, 0., Griffin, D. E. & Chase, J. (1977) Science 197, 376-378.
59. Payne, S., Parekh, B., Montelaro, R. C. & Issei, C. J. (1984)J. Gen. Virol. 65, 1395-1399.
60. Narayan, O., Sheffer, D., Griffin, D. E., Clements, J. & Hess, J. (1984) J. Virol. 49, 349-355.
61. Hahn, B. H., Gonda, M. A., Shaw, G. M., Popovic, M., Hoxie, J. A., Gallo, R. C. & Wong-Staal, F. (1985) Proc. Natl. Acad. Sci. USA 82, 4813-4817.
62. Wong-Staal, F., Shaw, G. M., Hahn, B. H., Salahuddin, S. Z., Popovic, M., Markham, P., Redfield, R. & Gallo, R. C. (1985) Science 229, 759-762.
63. Robert-Guroff, M., Brown, M. & Gallo, R. C. (1985) Nature (London) 316, 72-74.
64. Weiss, R. A., Clapham, P. R., Cheingsong-Popov, R., Dalgleish, A. G., Came, C. A., Weller, I. V. D. & Tedder, R. S. (1985) Nature (London) 316, 69-72.
________________
[i] Laboratory of Cell and Molecular Structure. Program Resources, Inc., National Cancer Institute-Frederick Cancer Research Facility, Frederick, MD 21701
[ii] Departments of Neurology and Molecular Biology and Genetics, Johns Hopkins University School of Medicine, Baltimore, MD 21205
[iii] Laboratory of Tumor Cell Biology, National Cancer Institute, Bethesda, MD 20892
[iv] Program Resources. Inc., National Cancer Institute-Frederick Cancer Research Facility, Frederick, MD 21701
Communicated by Paul C. Zamecnik, January 2, 1986
The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked" advertisement" in accordance with 18 U.S.C. §1734 solely to indicate this fact.
Abbreviations: AIDS, acquired immune deficiency syndrome; HTLV-I, -II, and -III, human T-cell lymphotropic virus type I, II, and III; CAEV, caprine arthritis encephalitis virus; EIAV, equine infectious anemic virus; tm, melting temperature; LA V, lymphadenopathy virus; Mo-MuLV, Moloney murine leukemia virus; RSV, Rous sarcoma virus; ORF, open reading frame; LTR, long terminal repeat.






